DrosGB (Drosophila Genomics Browser)
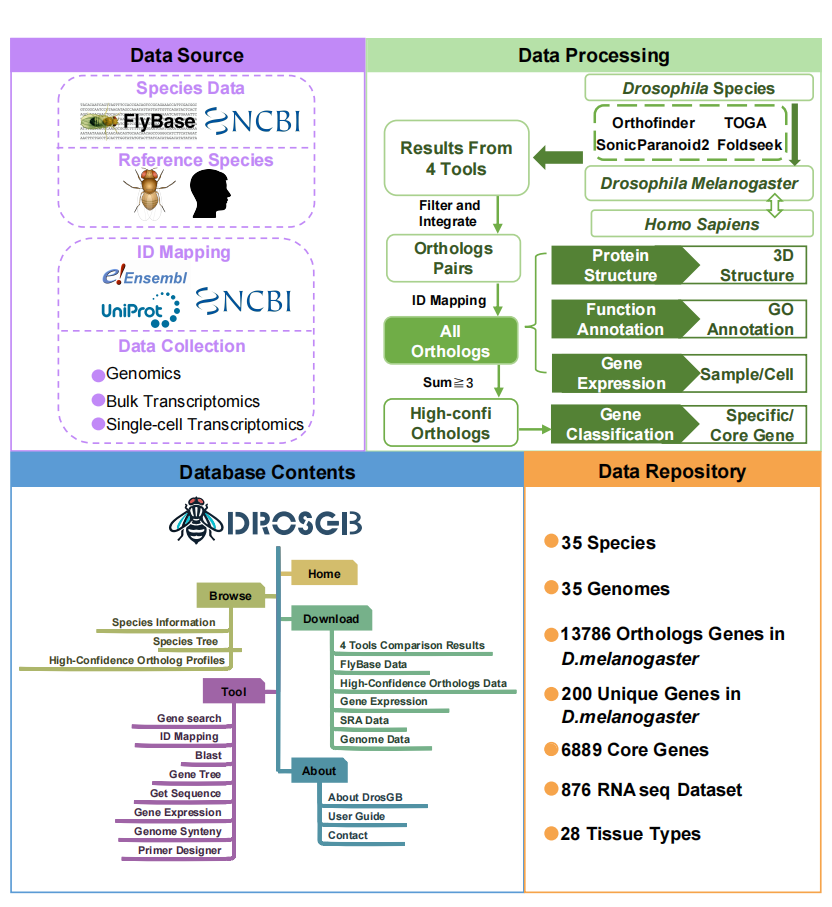
DrosGB is a comprehensive gene database focused on Drosophila species, designed to provide a one-stop data service and analysis platform for gene function research and comparative genomics across multiple species. DrosGB incorporates multi-omics data from 35 Drosophila species, including high-quality genome annotations, transcriptomic expression profiles, orthologous gene predictions, 3D protein structures, GO functional annotations, and single-cell transcriptomic maps.
The database integrates results from multiple mainstream orthology inference tools, such as OrthoFinder, SonicParanoid, Foldseek, and TOGA, and offers functional modules including gene ID search, rapid ortholog mapping, and BLAST alignment, facilitating the exploration of gene evolutionary relationships and functional characteristics.
Featured Tools
External Resources
Latest News & Updates
-
June 2025DrosGB Version 1.0 Released
-
October 2025DrosGB Version 2.0 ReleasedWe have added a 3D structure display of genes, as well as a number of new functions such as sequence search.
-
January 2026DrosGB Version 3.0 ReleasedWe have incorporated data from 15 additional Drosophila species.
How to Cite DrosGB
Zheng Q, Zhang C, Zhang J, et al. DrosGB: An integrated multi-omics database for comparative genomics and functional annotation of 35 Drosophila species. hLife 2026. https://doi.org/10.1016/j.hlife.2026.02.003.





